- Joined
- Jun 22, 2011
- Messages
- 2,431
- Points
- 83
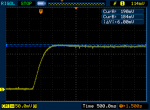
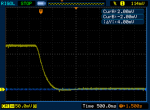
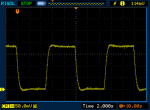
FWIW my Ophir also does the turn-on overshoot and sometimes the turn-off overshoot. I'm not sure if that's due to me moving the lasers while pressing the button. Might be, since any small movement will create that "ringing" effect on the graph.
Anyone with one of those remote wired tail caps?
Would also be interesting to see if this happens on other thermopile LPMs out there..
Anyone with one of those remote wired tail caps?
Would also be interesting to see if this happens on other thermopile LPMs out there..